B1.1: Complex RNA folding reactions populating various intermediates
B1.1: Complex RNA folding reactions populating various intermediates
This PhD student will investigate a ribozyme which catalyzes two distinct reactions. The sequence is a 79mer RNA able to adopt either a conformation exhibiting cleavage reactivity or a second conformation with ligase activity. Incorporation of photo-protecting groups at selected sites will force the sequence to adopt the ligase conformation. Light NMR experiments will be conducted, utilizing a laser coupled to the NMR spectrometer to release the photo-protecting groups while conformational switch can proceed. Since the desired RNA is too long to be synthesized by solid-phase synthesis in high yields, solid-phase synthesis has to be combined with enzymatic methods. The photo-protected part of the RNA will be synthesized by solid-phase synthesis in the Heckel group and the unmodified part will be synthesized enzymatically. The full-length photo-protected RNA will be obtained after enzymatic ligation and can be investigated by NMR spectroscopy.
Another project will be the development of a method for rapid preparation and screening of RNA secondary structures.
In addition, an approach for chemo-enzymatic synthesis of RNAs labelled with photo-protecting groups will be established. The Heckel group is involved in this project and will provide the photo-protected nucleoside 3',5'-bisphosphates. These nucleotides will be incorporated at an internal position of an RNA over our choice by a three-step enzymatic procedure.
 B1.2: Investigation of co-translational protein folding
B1.2: Investigation of co-translational protein folding
The majority of proteins starts to fold, while they are being synthesized by the ribosome. This process, called co-translational folding, should be studied using so called ribosome nascent chain complexes. Therefore, gamma-B crystalline, a bovine two-domain protein, which is known to fold in a co-translational manner, will be attached to the ribosome using the arrest peptide from the secretion monitor (SecM) of E. coli. Different lengths of the nascent chain, which are attached to the ribosome will provide snapshots of the translation. These complexes will be studied by solid state NMR enhanced by DNP and Cryo-EM. The released nascent chain will be studied by liquid state NMR. Using solid state NMR the folding of the nascent protein peptide chain can be measured inside and outside the ribosomal tunnel, while Cryo-EM provides a structure of the protein within the ribosomal tunnel.
Using all these techniques we can address different questions:
- Is disulfide bond formation possible inside the ribosomal tunnel?
- If yes, are those native disulfide bonds?
- Are possible disulfide bonds stable after they emerge from the ribosome?
- Is the structure of the protein within or at the ribosome the same as the one obtained after release?
The solid state NMR measurements are carried out with the help of the Glaubitz group and the Cryo-EM measurement with the help of the Frangakis group (Frankfurt).
B1.3: Complex strategies of light-regulation of oligonucleotides
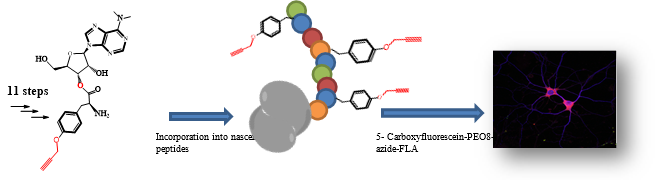
The aim of this PhD thesis is to design and synthesize new photocaged derivatives of puromycin for the spatiotemporal monitoring of newly translated proteins in living neurons. . Puromycin is an amino nucleoside antibiotic that mimics the 3´-end of Tyr-tRNA. It inhibits the protein biosynthesis during translation. This inhibition results in the release of the puromycylated nascent peptide chain. Such nascent chain can be detected utilizing an anti-puromycin antibody. Puromycin binds specifically at the C-terminus of the synthesized polypeptide chain and can thus be used as indicator of newly translated protein. A1.3 (Lisa M. Herzig, Wachtveitl group) will conducte biophysical experiments to characterize the photophysical properties of the compound.
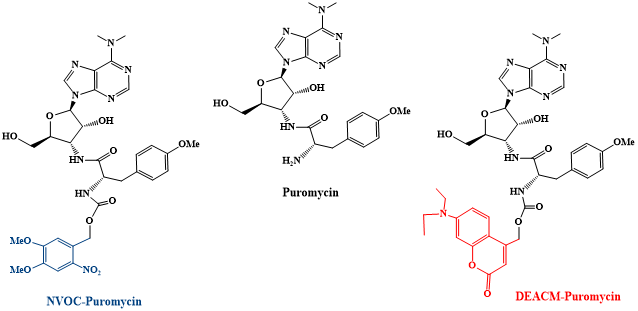
Fig.1: Photocaged DEACM- and NVOCpuromycin
Furthermore, during my PhD thesis I want to synthesize an alkyne analogue of puromycin that allows a more efficient incorporation of labeled amino acid into new translated proteins. The modified puromycin consists a ribonucleoside instead of aminonucleoside, which suppresses the inhibitory effect of puromycin. Nevertheless, the lower part containing an alkyne rest can bind to the newly synthesized nascent peptide chain with high incorporation rates.
Fig.2: Imaging protein synthesis in cells with an alkyne analog of puromycin
B1.4: Structural and dynamical investigation of RNA and DNA by NMR spectroscopy
The aim of the project is to understand better the structural effect of a single photolabile group on the DNA/RNA duplex stabilization. Using combined tools of NMR spectroscopic investigation of new NPE derivates (synthesized by PhD thesis A2.3), MD simulations (in collaboration with Hummer group – MPI Biophysics) and chemical screening with click chemistry (PhD thesis A2.4). This allows new generation of more efficiently destabilizing photolabile groups to be developed for biochemical applications.
With the help of NMR spectroscopy three dimensional structural models based on NOE constraints will be estabilished. These models will allow deeper understanding of the effect of configuration and steric effects of the photo cages. Furthermore these models and NMR data will be used to validate MD simulation of DNA double helix with attached photolabile group.
Questions to be answered:
- What is the effect of the configuration of the photo-cages on the DNA stability?
- How does a sterically more demanding substituent influences the destabilization property of the photo-cage?
- What modifications, interactions would allow maximum destabilization of a duble helix by a single photolabile group?
Suggested literature:
B1.5: Conformational selection in DNA: G-quadruplexes
Beside double-helix conformations, oligonucleotides can also form non-canonical structures including tetraplex structures formed by pairing of four strands. These tetraplex structures include G-rich G-quadrupluexes and C-rich i-motifs. These structures can act as regulatory units in cells. The project B1.5 will utilize the concept of conformational caging to characterize the kinetics of folding of these DNA regulatory units, following previous work in the group.
The work will involve the synthesis of caged oligonucleotide strands in the Heckel group, the ligation of caged and uncaged strands in the Schwalbe group and the time-resolved characterization of folding reactions by NMR spectroscopy.
Suggested literature:
B1.6: A two-function ribozyme and synthesis of large RNAs containing photocages
RNA molecules are unique in their ability to adopt multiple long-lived conformations with almost identical stability but high energetic separation. This can lead the population of multiple states for a single conformation that can exhibit distinctly different biological function. The investigation of their interconversion is particularly interesting. The project B1.6 will utilize the concept of conformational caging to characterize the kinetics of folding of bistable RNA structures.
A second aim of the project is to develop methods for the time-resolved footprinting of RNA conformation. Close collaboration with the Heckel and the Wachtveitl group are foreseen.
 B1.7: Synthesis of riboswitch-binding metabolites containing pho-tolabile protecting groups for biophysical studies
B1.7: Synthesis of riboswitch-binding metabolites containing pho-tolabile protecting groups for biophysical studies
Riboswitches represent an important class of RNA regulation elements. Upon of metobolites, ligands of low molecular weight, the expression of genes that are involved in the metabolism of the cognate ligand, are up- or downregulated. But to now, 25 different classes of riboswitches have been reported. In previous work, we have developed a caged version of one specific metabolite.
The project B1.7 will develop new caging strategy, ultimately to develop a chemical biology tool box of light regulatable small molecules.
Close collaboration with the Heckel and the Wachtveitl group are foreseen.
Suggested literature: