 B3.1: Time-resolved Solid-state NMR in Membrane Proteins in Lipid Cubic Phases
B3.1: Time-resolved Solid-state NMR in Membrane Proteins in Lipid Cubic Phases
The PhD student of this research area will be qualified in connected areas of expertise: He/She will be trained in solid-state NMR and in particular in fast detection techniques, in membrane protein biochemistry, in optical methods required to control light-triggered reactions and in preparing and handling of photo-labile compounds. The combination of these areas of expertise brings them into the unique position to develop novel approaches for observing kinetic events within biological membranes.
The PhD project involves developing an approach in which biochemical events mediated by membrane proteins can be triggered by light and followed in a time-resolved fashion by solid-state NMR. Solid-state NMR and in particular MAS-NMR is the method of choice for membrane protein studies directly within the lipid bilayer. However, time-resolved studies are challenging due to limited sensitivity and novel approaches for light triggering have to be developed. The problem of sensitivity can be solved by fast detection schemes improving signal-to-noise per time and by the use of dynamic nuclear polarization. For light triggering, problems with optical density and mixing in liposome preparations have to be overcome. A solution to these problems is given by using lipid cubic phases (LCPs). In these phases, lipid bilayers, usually formed by monoolein, follow an infinite periodic minimal surface, which divides the space into networks of water channels. In contrast to liposomes, LCPs are optically transparent and all binding sites of embedded membrane proteins remain fully accessible. Therefore, the mixing step could be optimized by using caged compounds, which are activated within the LCP inside the MAS rotor.
This approach will be established for the membrane protein diacylgylcerol kinase (DAGK). DAGK phosphorylates diacylglycerol within the membrane through the transfer of the gamma-phosphate from ATP. The full enzymatic reaction can be detected conveniently by 31P-MAS NMR. The reaction will be triggered by the release of caged ATP, caged Mg2+ or by activating caged diacylglycerol.
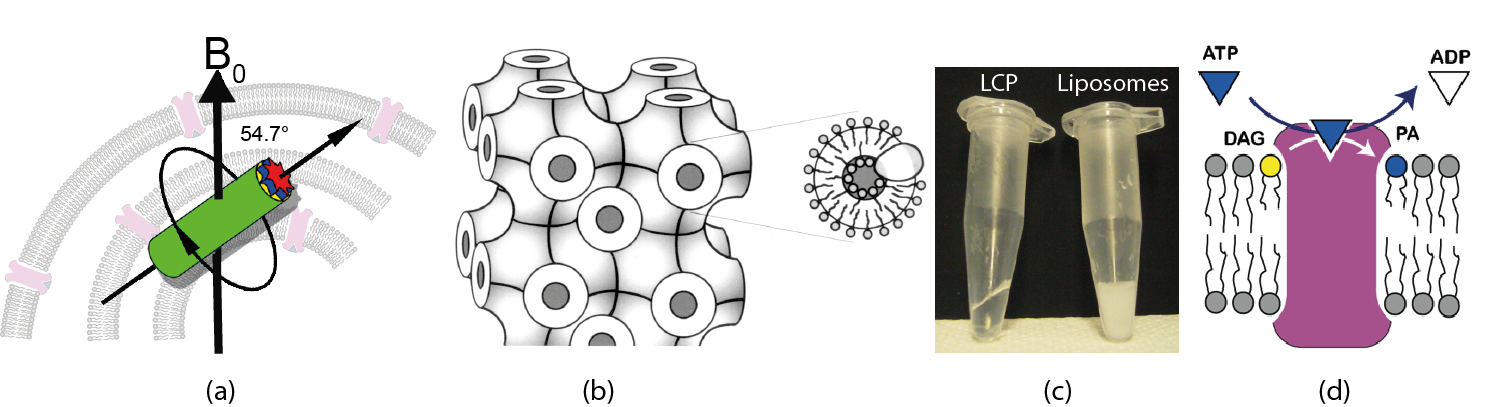
MAS-NMR is usually performed on pellets of proteolipsomes (a). Membrane proteins can be also incorporated into lipid cubic phases (LCP), in which bilayers follow an infinite periodic minimal surface dividing the space into a network of water channels (b). Comparison of LCP and liposome samples containing the membrane protein DAGK (c). DAGK is a membrane embedded enzyme, which phosphorylates diacylglycerol at the expense of ATP (d).
Suggested literature:
- "Perspectives in Enzymology of Membrane Proteins by Solid-State NMR", S. J. Ullrich, C. Glaubitz, Acc. Chem. Res. 2013, 46, 2164-2171.
- "Interfacial enzyme kinetics of a membrane bound kinase analyzed by real-time MAS-NMR", S. J. Ullrich, U. A. Hellmich, S. Ullrich, C. Glaubitz, Nat. Chem. Biol. 2011, 7, 263-270
B3.2: Development and application of light-controlled tools for MAS-NMR studies of membrane proteins
This project aims to develop novel techniques by which some parameters of liposomal environment, such as membrane lateral pressure, pH gradient, substrate release, etc., can be stimulated by light under nonequilibrium condition. This approach enables us to shed light on the transient conformations of membrane proteins and hence, on their complex functions within lipid membrane. Ethidium multidrug resistance protein (EmrE) is a small membrane transporter which antiports polyaromatic drugs in exchange of proton. As one part of this project, caged-protons are employed to be light-irradiated to establish reversible pH gradients throughout liposome environment. A variety of Solid-state NMR spectroscopy techniques (ssNMR) are then exploited to investigate the effects of generated pH gradients on the transient conformations of EmrE. Diacylglycerol kinase (DgkA) is the second membrane protein also interested as part of this project. In Escherichia coli, DgkA is responsible for the phosphorylation of diacylglycerol to phosphatidic acid in expense of triphosphate. SSNMR and time-resolved optical spectroscopy are the techniques of choice by which the impacts of lightswitchable lipids are studied on the conformation/function of DgkA, and on the mechanical-chemical properties of lipid membranes

